OCR Improvements: An Early Analysis
Optical character recognition (OCR) plays a critical part in BHL’s contributions to the scientific community. OCR in and of itself is a remarkable achievement, converting images of typewritten text to computer-readable text with “pretty good” accuracy. OCR on handwritten text is an even greater challenge to address and is beyond the scope of the improvements discussed here. The scientific work that BHL supports demands the best accuracy that we can provide using available tools, and let’s be honest, available budgets.
Recently, our colleagues at the Internet Archive made the transition away from the ABBYY FineReader OCR software to the Tesseract Open Source OCR engine. Over the past year or more, the OCR team at the Internet Archive has adapted and fine-tuned Tesseract to their workflows. Our first impression is that Tesseract OCR is more than “pretty good” in its ability to identify text from the page images provided to it.
The downside to this is that the Internet Archive has rightfully chosen to not re-process all existing text content through the Tesseract OCR engine. This is a prohibitively expensive and time-consuming prospect given that they have 35 million text-based items and reprocessing them would take several years and use up resources that could otherwise be used for gathering new content.
However, in the interests of supporting the efforts of the BHL community, the BHL Tech Team is working with our Internet Archive partner to reprocess some of BHL’s oldest content with the newest available version of Tesseract OCR. We are currently in a testing phase, and this blog post details some of our early results.
Selection
The first step in the process was to identify which versions of which OCR engine was used on BHL’s content. This was a simple matter of checking the “OCR” metadata value at the Internet Archive for each of the 263,000 items at BHL. The results as of December 2020 were:
| “OCR” Metadata Value | Count |
|---|---|
| ABBYY FineReader 8.0 | 80,447 |
| ABBYY FineReader 9.0 | 35,276 |
| ABBYY FineReader 11.0 | 45,679 |
| ABBYY FineReader 11.0 (Extended OCR) | 49,086 |
| Tesseract 4.1.1 | 83 |
| No OCR value | 39,530 |
We suspect that those items with No OCR value are the very oldest content at BHL and were processed with an unknown OCR engine, or ABBYY FineReader 8.0 or earlier.
For a test and ultimately for moving forward with reprocessing, we chose the 80,000 items with ABBYY FineReader 8.0. We will likely add the other 39,500 items with No OCR value.
The Process
The steps to reprocess the OCR for an item are simple. We delete a few critical files at the Internet Archive and issue a “derive” command to restore them. In doing so, the OCR is regenerated and the “OCR” metadata value is updated accordingly.
The challenging part of the process is time and resources. OCR is a computationally expensive process and it can take dozens of minutes to several hours to create the OCR for a single Item. We also must be aware that we can potentially take computing resources from other activities at the Internet Archive, so we issue the “derive” command at a lower priority than other Internet Archive activities, effectively using up spare or unused resources as they are available.
In other words, we’re being good citizens of the Internet Archive ecosystem.
The Results
Since this is what you came for, in summary, the results are very good and this is a worthwhile effort.
Our tests for evaluating the results are a combination of visual inspection and computational analysis. For backup and local analysis purposes, BHL keeps a copy of all content at the Internet Archive, but its updates are currently disabled during this testing phase. Using this backup copy as the source of “old” content, we can compare it to the “new” content at the Internet Archive.
Analysis 1: Misspellings
The first analysis is to simply check for misspelled words. While at first glance this seems too simplistic, OCR operates on a letter-by-letter analysis of the text and often has challenges correctly identifying a letter and will often produce invalid words.
Using the Linux aspell tool, we can find misspelled words and count them using a command such as the following:
cat FILENAME | aspell list | wc -l
This command sends the contents of FILENAME to aspell, which lists misspelled words, one per line, then sends that list to the word-count wc command to list the number of lines. Using a test case of 1955seventyoneye1955harr, we count the misspellings for the old and new versions:
- Old OCR Text (from 2013): 2,357 misspelled words
- New OCR Text (from 2020): 1,639 misspelled words
There are expected commonalities between the two lists of misspelled words. Proper or scientific names such as Elberta and Harrison or varietal names such as Dixired or Redhaven don’t exist in the standard English dictionary. Additionally, legitimate misspellings in the printed text appear, such as Recomemnded, appear in the results. Regardless, the greatly reduced number of misspellings is a good indicator of OCR accuracy.
Analysis 2: Visual Inspection
One of the most important downstream effects of improved OCR is improved identification of scientific names. BHL partners with the Global Names Architecture (GNA) to identify scientific names in the BHL text. Focusing on this, we can see that improvements in the OCR reveal more scientific names that were missed in the past.
Example A: Ligustrum ovalifolium
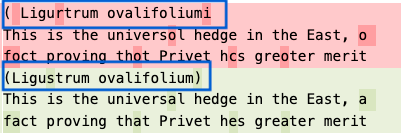
The original page image at BHL discusses the California Privet (Ligustrum ovalifolium).
The OCR comparison of the old engine (in red) and the new (in green) shows that the new Tesseract OCR engine was better able to convert the words to text. This will ultimately cause this name to appear in BHL’s list of scientific names on this page of the document where currently it may not appear.
Example B: Multiple names
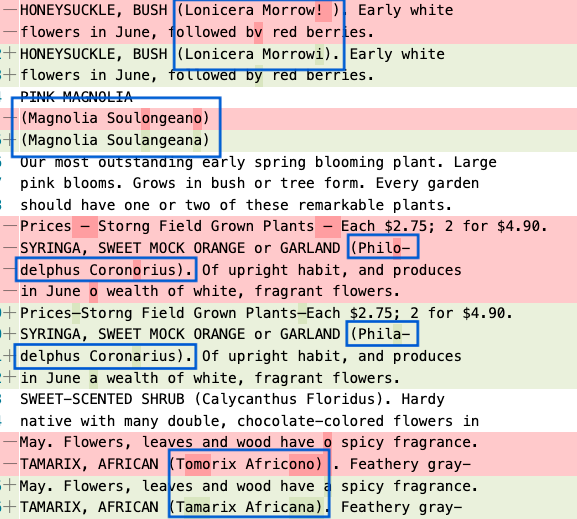
A second example showing numerous taxon names that are now correctly identified by the OCR. It’s worth mentioning that this page in BHL includes the genus (Magnolia or Lonicera) but not the full species name. We expect that the full name will appear in BHL with the improved OCR.
Analysis 3: Scientific Name Finding
As mentioned earlier, BHL partners with the Global Names Architecture (GNA) to identify scientific names in the OCR content of BHL. While we use APIs to perform this function, GNA also offers a command line tool to process a body of text to identify scientific names.
Similar to counting the misspelled words, we use a series of Linux commands to process the OCR text and count the scientific names found in the text.
gnfinder find FILENAME -c -s 1,3,4,9,11,12,167,172,179,181 | jq '.names[] .verification.BestResult.matchedName' | sort | uniq | wc -l
gnfinder is the command to find the scientific names. This command returns JSON content, which we then send to the jq command to count the number of BestResult.matchedName elements in the JSON. Then we sort, get the unique names, and count them with wc. A sample of the JSON output looks like:
{
"type": "Uninomial",
"verbatim": "(Lonicera",
"name": "Lonicera",
"odds": 93678.22872366496,
"annotation": "",
"verification": {
"BestResult": {
"dataSourceId": 1,
"dataSourceTitle": "Catalogue of Life",
"taxonId": "4091239",
"matchedName": "Lonicera",
"matchedCanonical": "Lonicera",
"currentName": "Lonicera",
"classificationPath":
"Plantae|Tracheophyta|Magnoliopsida|Dipsacales|
Caprifoliaceae|Lonicera",
"classificationRank":
"kingdom|phylum|class|order|family|genus",
"classificationIDs":
"3939764|3942634|3942724|3942969|3942971|4091239",
"matchType": "ExactMatch"
}
},
[...]
}
Counting these for our example 1955seventyoneye1955harr, we find:
- Old OCR Scientific Names: 20 unique names found
- New OCR Scientific Names: 38 unique names found
This is a simple case indicating that an additional 18 unique names were found in the content. Taking a random sample of 10 other items shows the following larger differences in the number of unique scientific names found:
| Item Identifier | Unique Names Found in OCR | |
|---|---|---|
| Old | New | |
| guidebooksofexcu03inte | 190 | 190 |
| dissectionofdoga00howe | 23 | 25 |
| ueberliasbeta00schl | 38 | 55 |
| mobot31753003413330 | 641 | 920 |
| CUbiodiversity1249031-9750 | 1,042 | 1,244 |
| annalesdelasoci2627188283soci | 2,786 | 3,046 |
| weiterebeobachtu00kl | 59 | 62 |
| verhandlungender42zool | 2,487 | 2,806 |
| etudedesfleu1865cari | 1,077 | 1,148 |
| bulletinbiologiq47univ | 750 | 1,127 |
It is worth noting that a visual inspection of these names indicates there is some fine-tuning remaining. gnfinder identifies both binomial names (genus and species) and uninomial names (genus only) in the content. There look to be instances where names are found in the new OCR that don’t exist in the content, but are incorrectly being identified as uninomials. gnfinder provides a type of score that must be fine tuned for the new OCR in order to reduce this effect. This is a task for future discussion.
Summary
While there is further work to do in loading this new content into BHL and in the scientific-name-finding part of the process, these initial results are encouraging and are enough to help us make the decision to continue reprocessing the OCR using the Internet Archive’s installation of Tesseract OCR.
This journey is years in length. Even if we were to process at the highest priority (something we would never consider), we are planning to affect nearly half of BHL’s 263,000 items. Our current rate of progress at the aforementioned lower priority is approximately 100 items per day. At such a pace, the 120,000 items will take three years to complete.
Future blog posts will occur as there are more updates to share with the BHL community.





Leave a Comment